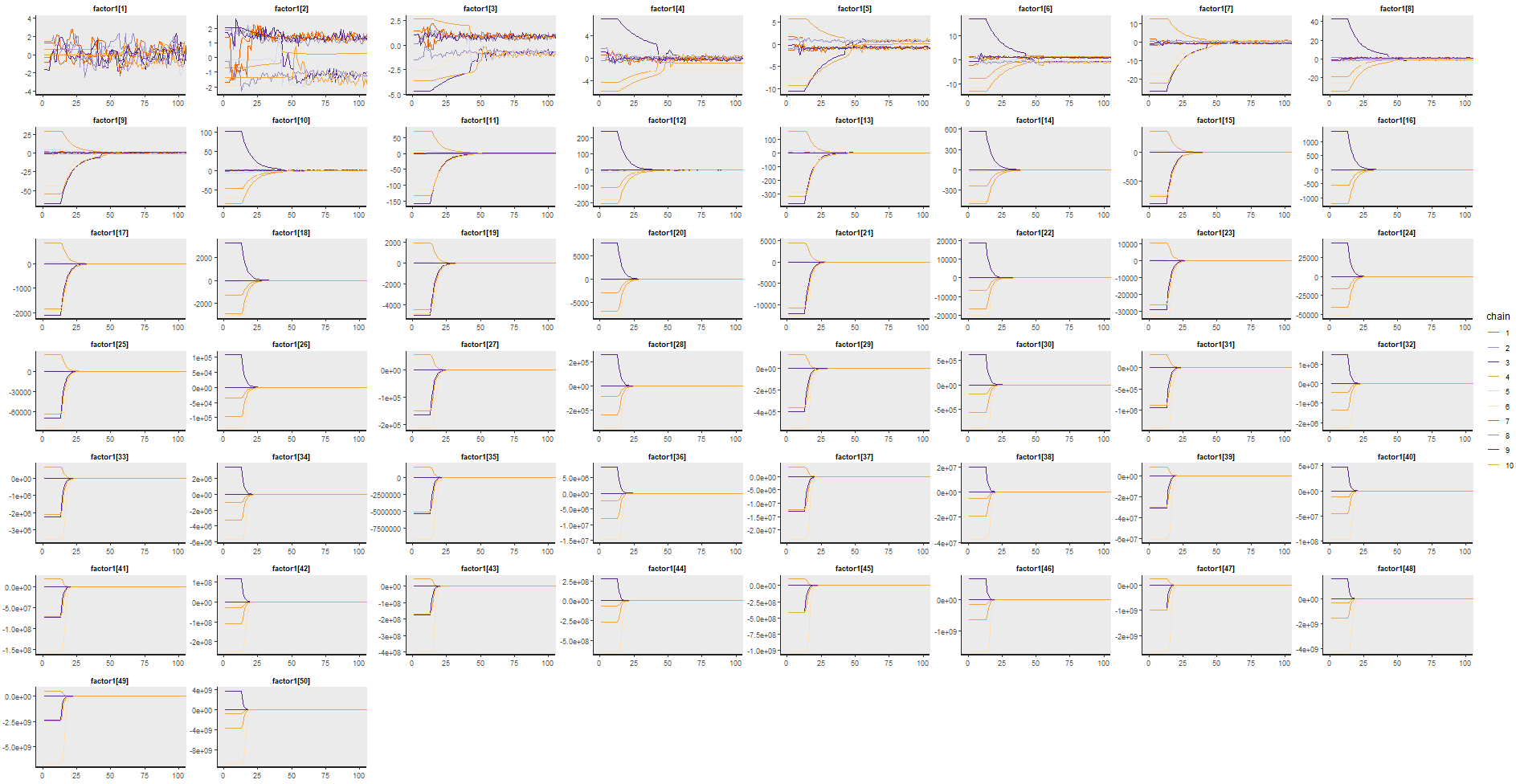
I played around with the code and tried a few things. One thing that seems to help to me is to constrain the factors to be stationary or at most random walk, by imposing them to be between -1 and 1. That should help rule out the more explosive behaviour, and I feel it’s not an unreasonable restriction in this type of setting.
matrix<lower=-1,upper=1>[K, K] ar_facts;
This already imposes a stronger regularization than the normal(0, 10) prior you already have, but you may want to tweak that prior further.
With this approach, I generally get good behaviour, although indeed a couple of misbehaved chains seem to have the top loads close to 0 and get stuck there. Then the remaining variables get the wrong side in their loadings, and thus the estimated factors also have the wrong sign, but “correct” with a sign switch. I tried to ensure I was running the model with strong loadings on the first variable to ensure identifiability, as otherwise it’s easier for that loading to be estimated as small positive 0 and then the remaining variables to end up in different chains on different sides of positive and negative. And I also played with simulating data with ar_facts with a larger number (0.7), as I wondered if -0.1 is too weak for any discernible factor structure to be estimated. Still, this did not seem to be enough to get good results every time I ran with your example (N = 5, T = 300).
This is a strange conjecture, and maybe I can’t explain it clearly. I think the main issue is the top load getting stuck at 0, and the HMC being unable to move away from that. I wonder if what’s happenning is the following. The top load must be positive due to the cholesky_factor_cov constraint you impose, I think. And in simulations, I ensured that it was strongly positive (among the largest absolute values). Suppose that the estimation of the top load starts close to 0. What may happen as the DFM grows is that the model is still able to have a factor that is driving all the other N-1 variables reasonably well, but with the opposite sign, and will just ignore that first variable, imagining it to have a very small loading close to 0. As this has still a high likelihood (as the single factor can explain N-1 variables), it’s hard for it to move away from 0, as that would require a large move if the first_load is high and would require a sign switch for all the other loadings and the factors. An additional signal in this direction is that if I simulate data with all positive factors AND I restrict bottom_loads to be positive, the estimation avoids the R-Hat and E-BFMI issues (although in my only attempt it still had a small number of divergent transitions). However, in practice this may be hard to ensure, or would require you to do some pre-processing that may not even be obvious how to do.
I hope these are useful clues. If you learn anything about the topic, please keep posting, as I’m also interested in estimating these models in Stan.