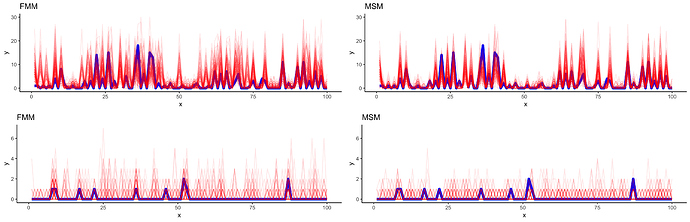
I want to implement a zero-inflated negative binomial model for longitudinal data (assume a balanced panel that is zero inflated where non-zero observations are positive counts that are negative binomial distributed) with a Markov-Switching process. I.e. instead of assuming that temporally adjacent states are independent I assume that they are informative about immediately adjacent states as in the DGP the discrete process that determines whether we see a 0 or a draw from a negative binomial is a first order Markov process.
I’ve two versions at this point though neither can recover the draws from the prior distributions and the distributions that are being recovered usually are far away from the prior draw. A specific ``problem’’ here is that I know the state with certainty when I observe counts, i.e. Pr(S_t = NB \vert y_{1:t}) = 1 and Pr(S_t = 0process \vert y_{1:t}) = 0 but I don’t know how to implement this correctly while also adding the correct density to the posterior. So one implementation ignores p(S_t\vert y_{1:t - 1}) by considering it as equal to 1 for NB and 0 for the 0 process. The other always adds the density for p(S_t\vert y_{1:t - 1}) but gives the other state a very small probability i.e. log(0.0000000001). Yet, that doesn’t represent the DGP accurately and feels like a bad workaround (which also doesn’t work).
I would greatly appreciate any ideas you might have.
R
dgp_zip_msm_tvtp_v5 <- function(N, T, Mz, Mx){
# startstop
Tn <- rep(T, N)
if (N == 1){
start.stop <- array(c(1, T), dim = c(1, 2))
} else {
start.stop <- cbind(c(1, cumsum(Tn[1:(N - 1)]) + 1), Tn)
}
NT <- sum(Tn)
# predictors
Z <- matrix(rnorm(NT * Mz), ncol = Mz, nrow = NT)
X <- matrix(rnorm(NT * Mx), ncol = Mx, nrow = NT)
# parameters
lambda <- rnorm(Mx, 0, 1) # nb parameters
zeta <- matrix(rnorm(Mz * 2), ncol = 2, nrow = Mz) # discrete process
theta <- rnorm(1, 40, 10) # dispersion
# TP
A <- array(NA, dim = c(NT, 2, 2))
for (nt in 1:NT){
A[nt, 1, 1] = pnorm(Z[nt, ] %*% zeta[, 1])
A[nt, 2, 2] = pnorm(Z[nt, ] %*% zeta[, 2])
A[nt, 1, 2] = 1 - A[nt, 1, 1]
A[nt, 2, 1] = 1 - A[nt, 2, 2]
}
s <- rep(NA, times = NT)
K.seq <- seq(1, 2)
for (n in 1:N){
t0 = start.stop[n, 1]
s[t0] <- t(rmultinom(1, 1, prob = rep(0.5, 2))) %*% K.seq
for (t in 2:T){
t0 = start.stop[n, 1] + t - 1
t1 = start.stop[n, 1] + t - 2
s[t0] <- t(rmultinom(1, 1, prob = A[t0, s[t1], ])) %*% K.seq
}
}
y <- rep(NA, times = NT)
for (n in 1:N){
for (t in 1:T){
t0 <- start.stop[n, 1] + t - 1
if (s[t0] == 1){
y[t0] <- 0
} else {
y[t0] <- rnbinom(1, size = theta, mu = exp(X[t0, ] %*% lambda))
}
}
}
stan_data <- list(N = N, T = T, NT = NT, y = y, Mz = Mz,
Mx = Mx, Z = Z, X = X, startstop = start.stop)
parameters <- list(A = A, lambda = lambda, zeta = zeta, S = s, theta = theta)
return(list(stan_data = stan_data, parameters = parameters))
}
Stan version 1
data {
int<lower=0> N;
int<lower=0> NT;
int<lower=0> y[NT];
int startstop[N, 2];
// Predictors dimensions
int<lower = 0> Mz;
int<lower = 0> Mx;
//Predictors
matrix[NT, Mz] Z;
matrix[NT, Mx] X;
}
parameters {
simplex[2] pi1[N];
// discrete
matrix[Mz, 2] zeta;
// continuous
vector[Mx] lambda; //predictors
real<lower = 0> theta; // dispersion
}
transformed parameters {
vector[2] logalpha[NT];
simplex[2] A[NT, 2];
vector[2] logA[NT, 2];
vector[NT] lambda_;
for (nt in 1:NT){
A[nt, 1, 1] = normal_cdf( Z[nt] * zeta[:, 1], 0, 1);
A[nt, 2, 2] = normal_cdf( Z[nt] * zeta[:, 2], 0, 1);
A[nt, 1, 2] = 1 - A[nt, 1, 1];
A[nt, 2, 1] = 1 - A[nt, 2, 2];
}
logA = log(A);
lambda_ = exp(X * lambda);
for (n in 1:N){
// t = 1
{
int t0 = startstop[n, 1];
if (y[t0] > 0){
logalpha[t0, 2] = neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta);
logalpha[t0, 1] = 0.0;
} else if (y[t0] == 0) {
logalpha[t0, 2] = neg_binomial_2_lpmf(0 | lambda_[t0], theta) + log(pi1[n, 2]);
logalpha[t0, 1] = log(pi1[n, 1]);
}
}
// t = 2:T
for (t in 2:startstop[n, 2]){
{
int t0 = startstop[n, 1] + t - 1;
int t1 = startstop[n, 1] + t - 2;
if (y[t1] > 0){
if (y[t0] > 0){
logalpha[t0, 2] = neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta);
logalpha[t0, 1] = 0.0;
} else if (y[t0] == 0){
logalpha[t0, 2] = logA[t0, 2, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta);
logalpha[t0, 1] = logA[t0, 2, 1];
}
} else if (y[t1] == 0){
if (y[t0] > 0){
logalpha[t0, 2] = neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta);
logalpha[t0, 1] = 0.0;
} else if (y[t0] == 0){
logalpha[t0, 2] = log_sum_exp(
logA[t0, 1, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta) + logalpha[t1, 1],
logA[t0, 2, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta) + logalpha[t1, 2]);
logalpha[t0, 1] = log_sum_exp(
logA[t0, 1, 1] + logalpha[t1, 1],
logA[t0, 2, 1] + logalpha[t1, 2]);
}
}
}
}
}
}
model {
for (n in 1:N) target += dirichlet_lpdf(pi1[n] | rep_vector(1, 2));
for (m in 1:Mx) target += normal_lpdf(lambda[m] | 0, 3);
for (j in 1:2) target += normal_lpdf(zeta[j] | 0, 3);
target += normal_lpdf(theta | 0, 2);
for (nt in 1:NT){
if (y[nt] > 0){
target += logalpha[nt, 2];
} else {
target += logalpha[nt];
}
}
}
generated quantities {
vector[2] prS[NT];
// prS
for (nt in 1:NT){
if (y[nt] > 0){
prS[nt, 2] = 1; prS[nt, 1] = 0;
} else {
prS[nt] = softmax(logalpha[nt]);
}
}
}
Stan version 2
data {
int<lower=0> N;
int<lower=0> NT;
int<lower=0> y[NT];
int startstop[N, 2];
// Predictors dimensions
int<lower = 0> Mz;
int<lower = 0> Mx;
//Predictors
matrix[NT, Mz] Z;
matrix[NT, Mx] X;
}
transformed data {
real pointllh = 0.00000000001;
}
parameters {
simplex[2] pi1[N];
// discrete
matrix[Mz, 2] zeta;
// continuous
vector[Mx] lambda; //predictors
real<lower = 0> theta; // dispersion
}
transformed parameters {
vector[2] logalpha[NT];
simplex[2] A[NT, 2];
vector[2] logA[NT, 2];
vector[NT] lambda_;
for (nt in 1:NT){
A[nt, 1, 1] = normal_cdf( Z[nt] * zeta[:, 1], 0, 1);
A[nt, 2, 2] = normal_cdf( Z[nt] * zeta[:, 2], 0, 1);
A[nt, 1, 2] = 1 - A[nt, 1, 1];
A[nt, 2, 1] = 1 - A[nt, 2, 2];
}
logA = log(A);
lambda_ = exp(X * lambda);
for (n in 1:N){
// t = 1
{
int t0 = startstop[n, 1];
if (y[t0] > 0){
logalpha[t0, 2] = neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta);
logalpha[t0, 1] = log(pointllh);
} else if (y[t0] == 0) {
logalpha[t0, 2] = neg_binomial_2_lpmf(0 | lambda_[t0], theta) + log(pi1[n, 2]);
logalpha[t0, 1] = log(pi1[n, 1]);
}
}
// t = 2:T
for (t in 2:startstop[n, 2]){
{
int t0 = startstop[n, 1] + t - 1;
int t1 = startstop[n, 1] + t - 2;
if (y[t1] > 0){
if (y[t0] > 0){
logalpha[t0, 2] = log_sum_exp(
logA[t0, 2, 2] + neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta) + logalpha[t1, 2],
logA[t0, 1, 2] + neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta) + logalpha[t1, 1]);
logalpha[t0, 1] = log_sum_exp(
log(pointllh) + logalpha[t1, 2] + logA[t0, 2, 1],
log(pointllh) + logalpha[t1, 1] + logA[t0, 1, 1]);
} else if (y[t0] == 0){
logalpha[t0, 2] = logalpha[t1, 2] + logA[t0, 2, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta);
logalpha[t0, 1] = logalpha[t1, 2] + logA[t0, 2, 1] ;
}
} else if (y[t1] == 0){
if (y[t0] > 0){
logalpha[t0, 2] = log_sum_exp(
logalpha[t1, 1] + logA[t0, 1, 2] + neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta),
logalpha[t1, 2] + logA[t0, 2, 2] + neg_binomial_2_lpmf(y[t0] | lambda_[t0], theta));
logalpha[t0, 1] = log_sum_exp(
log(pointllh) + logalpha[t, 2] + logA[t0, 2, 1],
log(pointllh) + logalpha[t, 1] + logA[t0, 1, 1]);
} else if (y[t0] == 0){
logalpha[t0, 2] = log_sum_exp(
logA[t0, 1, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta) + logalpha[t1, 1],
logA[t0, 2, 2] + neg_binomial_2_lpmf(0 | lambda_[t0], theta) + logalpha[t1, 2]);
logalpha[t0, 1] = log_sum_exp(
logA[t0, 1, 1] + logalpha[t1, 1],
logA[t0, 2, 1] + logalpha[t1, 2]);
}
}
}
}
}
}
model {
for (n in 1:N) target += dirichlet_lpdf(pi1[n] | rep_vector(1, 2));
for (m in 1:Mx) target += normal_lpdf(lambda[m] | 0, 3);
for (j in 1:2) target += normal_lpdf(zeta[j] | 0, 3);
target += normal_lpdf(theta | 0, 2);
for (n in 1:N){
{
int end = startstop[n, 1] + startstop[n, 2] - 1;
target += logalpha[end];
}
}
}
generated quantities {
vector[2] prS[NT];
// prS
for (nt in 1:NT){
if (y[nt] > 0){
prS[nt, 2] = 1; prS[nt, 1] = 0;
} else {
prS[nt] = softmax(logalpha[nt]);
}
}
}