In the openstatsware group (formerly the ASA Biopharmaceutical Software Engineering Working Group), we are developing an R package to fit Bayesian mixed models of repeated measures (MMRMs): GitHub - openpharma/brms.mmrm: R package to run Bayesian MMRMs using {brms}. Our package uses brms in the backend, and its goal is to simplify MMRM modeling using in an interface tailored to the needs of clinical trial data analysis.
We are trying to validate our implementation using simulation-based calibration (SBC). Initially we used brms::brm(sample_prior = "only") to simulate from the prior, and the rank statistics all looked uniform as we had hoped. But we do not want to stop there because it is arguably circular to simulate using brms and then analyze with brms. For extra assurance, and to convince ourselves that we understand what brms is doing, we are developing custom R code to simulate from the prior. And that is where things are not going smoothly.
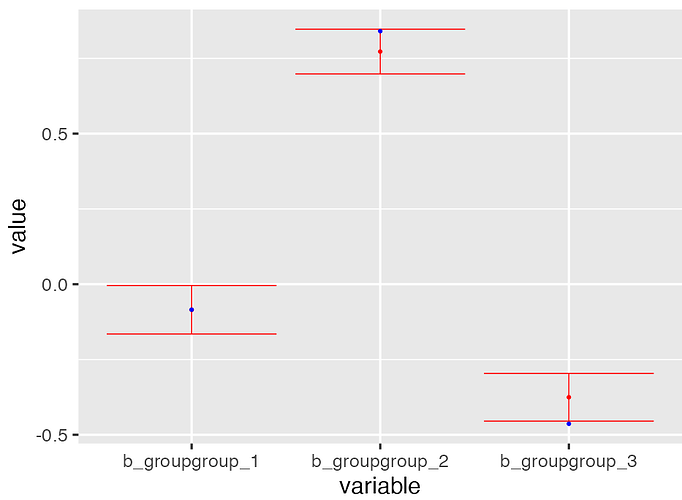
The reprex below runs an MMRM tailored to clinical trial data. We have independent patients who were randomized to 3 different treatment groups, and each patient is observed in 4 different discrete time points (scheduled study visits). The model formula uses treatment group levels as fixed effects (with no intercept), and the residuals within each patient have an unstructured (fully parameterized) correlation matrix. The correlation is modeled with a separation strategy and an LKJ prior.
brmsformula(
formula = response ~ 0 + group + unstr(time = time, gr = patient),
sigma ~ 0 + time
)
Oddly enough, the intercept-only equivalent appears well calibrated (response ~ unstr(time = time, gr = patient)), and so does a model with independent residuals (response ~ 0 + group). That alone makes me think the culprit is some sort of non-identifiability, but to me all the parameters seem like they should be easy to estimate.
Anyway, here is the code. The simulations unfortunately take a long time to run, especially for a reprex. I am using brms 2.20.4, rstan 2.32.3, and StanHeaders 2.26.28. See the very bottom for the rank statistics histograms and session info.
library(brms)
library(dplyr)
library(tibble)
library(tidyr)
#############
# FUNCTIONS #
#############
one_sbc_replication <- function(
chains = 4L,
warmup = 2000L,
iter = 4000L
) {
prior <- set_prior("lkj_corr_cholesky(1)", class = "Lcortime") +
set_prior("normal(0, 1)", class = "b", coef = "groupgroup_1") +
set_prior("normal(0, 1)", class = "b", coef = "groupgroup_2") +
set_prior("normal(0, 1)", class = "b", coef = "groupgroup_3") +
set_prior("normal(0, 1)", class = "b", dpar = "sigma", coef = "timetime_1") +
set_prior("normal(0, 1)", class = "b", dpar = "sigma", coef = "timetime_2") +
set_prior("normal(0, 1)", class = "b", dpar = "sigma", coef = "timetime_3") +
set_prior("normal(0, 1)", class = "b", dpar = "sigma", coef = "timetime_4")
formula <- brmsformula(
formula = response ~ 0 + group + unstr(time = time, gr = patient),
sigma ~ 0 + time
)
simulation <- simulate_data(formula = formula, prior = prior)
options(brms.backend = "rstan")
model <- brm(
data = simulation$data,
formula = formula,
prior = prior,
chains = chains,
cores = chains,
iter = iter,
warmup = warmup
)
get_sbc_ranks(model, simulation)
}
simulate_data <- function(formula, prior) {
n_group <- 3L
n_patient <- 100L
n_time <- 4L
patients <- tibble(
group = paste0("group_", rep(seq_len(n_group), each = n_patient)),
patient = paste0("patient_", seq_len(n_group * n_patient))
)
data <- expand_grid(patients, time = paste0("time_", seq_len(n_time)))
data$response <- 0
x <- make_standata(formula, data, prior = prior)$X
beta <- rnorm(n = n_group, mean = 0, sd = 1)
names(beta) <- paste0("b_", colnames(x))
b_sigma <- rnorm(n = n_time, mean = 0, sd = 1)
names(b_sigma) <- paste0("b_sigma_timetime_", seq_len(n_time))
sigma <- exp(b_sigma)
correlation <- trialr::rlkjcorr(n = 1L, K = n_time, eta = 1)
i <- rep(seq_len(n_time), each = n_time)
j <- rep(seq_len(n_time), times = n_time)
cortime <- as.numeric(correlation)[j > i]
names(cortime) <- sprintf("cortime__time_%s__time_%s", i[j > i], j[j > i])
covariance <- diag(sigma) %*% correlation %*% diag(sigma)
data <- data |>
mutate(mu = as.numeric(x %*% beta)) |>
mutate(index_patient = rep(seq_len(n_patient * n_group), each = n_time)) |>
group_by(index_patient) |>
mutate(response = MASS::mvrnorm(mu = mu, Sigma = covariance)) |>
ungroup() |>
select(-index_patient, -mu)
parameters <- c(beta, b_sigma, cortime)
stopifnot(!anyDuplicated(names(parameters)))
list(data = data, parameters = parameters)
}
get_sbc_ranks <- function(model, simulation) {
draws <- posterior::as_draws_matrix(model)
draws <- draws[, setdiff(colnames(draws), c("lprior", "lp__"))]
truth <- simulation$parameters
stopifnot(all(sort(names(truth)) == sort(colnames(draws))))
draws <- draws[, names(truth)]
ranks <- SBC::calculate_ranks_draws_matrix(variables = truth, dm = draws)
tibble::as_tibble(as.list(ranks))
}
##############
# SIMULATION #
##############
# I used an SGE cluster, so this is how I set up the crew controller:
controller <- crew.cluster::crew_controller_sge(
name = "brms-mmrm-sbc",
workers = 100L,
seconds_idle = 30,
seconds_launch = 1800,
launch_max = 3L,
script_lines = "module load R/4.2.2",
sge_cores = 4L
)
# But if you have different resources, you may want to choose
# a different crew launcher plugin, e.g.:
# controller <- crew::crew_controller_local()
# Run the simulations:
controller$start()
tasks <- controller$map(
command = one_sbc_replication(chains = 4L, warmup = 2000L, iter = 4000L),
iterate = list(index = seq_len(100L)),
globals = as.list(globalenv()),
packages = c("brms", "dplyr", "tibble", "tidyr")
)
controller$terminate()
simulations <- bind_rows(tasks$result)
###########
# RESULTS #
###########
library(tidyr)
results <- pivot_longer(
simulations,
cols = everything(),
names_to = "parameter",
values_to = "rank"
)
library(ggplot2)
plot <- ggplot(results) +
geom_histogram(
aes(x = rank),
breaks = seq(from = 0, to = max(results$rank), length.out = 10)
) +
facet_wrap(~parameter) +
theme_gray(16)
ggsave("plot.png", plot, width = 12, height = 10)
The rank statistic histograms in the plot below are far from uniform.
Session info:
R version 4.2.2 Patched (2022-11-30 r83413)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Red Hat Enterprise Linux
Matrix products: default
BLAS/LAPACK: /CENSORED/intel-2020/compilers_and_libraries_2020.0.166/linux/mkl/lib/intel64_lin/libmkl_gf_lp64.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ggplot2_3.4.0 tidyr_1.2.1 tibble_3.2.1 dplyr_1.1.3 brms_2.20.4
[6] Rcpp_1.0.9
loaded via a namespace (and not attached):
[1] nlme_3.1-160 matrixStats_0.63.0 xts_0.12.2
[4] threejs_0.3.3 rstan_2.32.3 tensorA_0.36.2
[7] tools_4.2.2 backports_1.4.1 utf8_1.2.3
[10] R6_2.5.1 DT_0.26 ggdist_3.2.0
[13] colorspace_2.0-3 withr_2.5.1 tidyselect_1.2.0
[16] gridExtra_2.3 prettyunits_1.1.1 processx_3.8.2
[19] Brobdingnag_1.2-9 emmeans_1.8.8 curl_4.3.3
[22] compiler_4.2.2 cli_3.6.1 binom_1.1-1.1
[25] arrayhelpers_1.1-0 shinyjs_2.1.0 sandwich_3.0-2
[28] colourpicker_1.2.0 posterior_1.4.1 scales_1.2.1
[31] dygraphs_1.1.1.6 checkmate_2.1.0 mvtnorm_1.1-3
[34] callr_3.7.3 QuickJSR_1.0.5 stringr_1.5.0
[37] digest_0.6.33 StanHeaders_2.26.28 base64enc_0.1-3
[40] pkgconfig_2.0.3 htmltools_0.5.4 fastmap_1.1.0
[43] htmlwidgets_1.5.4 rlang_1.1.1 shiny_1.7.3
[46] svUnit_1.0.6 farver_2.1.1 generics_0.1.3
[49] zoo_1.8-11 jsonlite_1.8.4 crosstalk_1.2.0
[52] gtools_3.9.4 distributional_0.3.1 inline_0.3.19
[55] magrittr_2.0.3 loo_2.5.1 bayesplot_1.10.0
[58] Matrix_1.5-3 munsell_0.5.0 fansi_1.0.5
[61] abind_1.4-5 lifecycle_1.0.3 stringi_1.7.8
[64] multcomp_1.4-20 MASS_7.3-58.1 pkgbuild_1.4.0
[67] plyr_1.8.8 grid_4.2.2 parallel_4.2.2
[70] promises_1.2.0.1 crayon_1.5.2 miniUI_0.1.1.1
[73] lattice_0.20-45 splines_4.2.2 SBC_0.2.0.9000
[76] ps_1.7.5 pillar_1.9.0 trialr_0.1.5
[79] igraph_1.5.1 markdown_1.7 estimability_1.4.1
[82] shinystan_2.6.0 reshape2_1.4.4 codetools_0.2-18
[85] stats4_4.2.2 rstantools_2.3.1.1 glue_1.6.2
[88] V8_4.2.2 RcppParallel_5.1.5 vctrs_0.6.4
[91] httpuv_1.6.6 purrr_1.0.2 gtable_0.3.1
[94] cachem_1.0.6 mime_0.12 xtable_1.8-4
[97] coda_0.19-4 later_1.3.0 survival_3.4-0
[100] shinythemes_1.2.0 memoise_2.0.1 tidybayes_3.0.2
[103] TH.data_1.1-1 ellipsis_0.3.2 bridgesampling_1.1-2