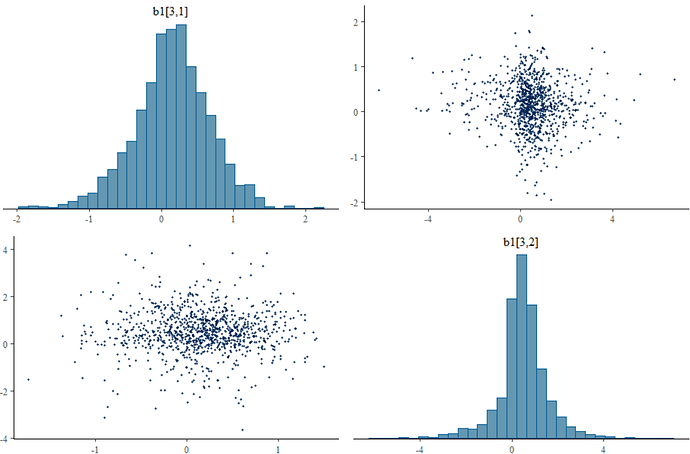
I am fitting a model for diagnostic test accuracy data. There are three tests and each can give three outcomes: positive, intermediate result, or negative. The model which assumes conditional dependence between tests works fine. However, when I try to model the dependence by introducting a random effect t into the probit regression model. I get a huge number of divergent transitions (~ 50% of the post-warmup iterations). Was wondering if anybody had any ideas.
Stan and R code below
data {
int<lower=2> nt; // number of tests
int<lower=0> N; // # of subjects
int<lower=1> res1[N]; // result of test 1
int<lower=1> res2[N]; // result of test 2
int<lower=1> res3[N]; // result of test 3
}
parameters {
vector[nt]<lower=0> a11_raw;
vector[nt] a10_raw;
vector<lower=0.0001>[nt] c11_raw;
vector<lower=0.0001>[nt] c10_raw;
vector[nt] b0_raw;
vector[nt] b1_raw;
vector[N] t; // subject-specific random effect
real<lower=0,upper=1> p; // disease prevalence
}
transformed parameters {
vector[nt] a11;
vector[nt] a10;
vector[nt] c11; // upper threshold for diseased (lower set to 0)
vector[nt] c10; // upper threshold for non-diseased
vector[nt] b0;
vector[nt] b1;
vector<lower=0,upper=1>[nt] Se;
vector<lower=0,upper=1>[nt] Sp;
vector<lower=0,upper=1>[nt] int_d;
vector<lower=0,upper=1>[nt] int_nd;
matrix<lower=0,upper=1>[N,nt] p1[2,3];
a11 = 2*a11_raw;
a10 = 2*a10_raw;
c11 = 2*c11_raw;
c10 = 2*c10_raw;
b0 = 2*b0_raw;
b1 = 2*b1_raw;
// overall / summary accuracy estimates, for each test
Se = 1-Phi_approx( (c11 - a11) ./ sqrt(1 + b1 .* b1) );
int_d = Phi_approx( (c11 - a11) ./ sqrt(1 + b1 .* b1) ) - Phi_approx( (- a11) ./ sqrt(1 + b1 .* b1) );
int_nd = Phi_approx( (c10 - a10) ./ sqrt(1 + b0 .* b0) ) - Phi_approx( (- a10) ./ sqrt(1 + b0 .* b0) );
Sp = Phi_approx( (- a10) ./ sqrt(1 + b0 .* b0) );
for (j in 1:nt) {
for (i in 1:N) {
// ordered probit regression model
// Non-diseased
p1[1,1,i,j] = Phi_approx( - a10[j] - t[i]*b0[j]) ; // prob of testing negative given non-diseased
p1[1,2,i,j] = Phi_approx(c10[j] - a10[j] - t[i]*b0[j]) - Phi_approx(- a10[j] - t[i]*b0[j]) ; // intermedtiate result given non-diseased
p1[1,3,i,j] = 1-Phi_approx(c10[j] - a10[j] - t[i]*b0[j]) ; // prob of testing positive given non-diseased
// diseased
p1[2,1,i,j] = Phi_approx( - a11[j] - t[i]*b1[j]) ;
p1[2,2,i,j] = Phi_approx(c11[j] - a11[j] - t[i]*b1[j] ) - Phi_approx( - a11[j] - t[i]*b1[j]) ;
p1[2,3,i,j] = 1-Phi_approx(c11[j] - a11[j] - t[i]*b1[j]) ;
}
}
}
model {
a11_raw ~ std_normal(); // ~ normal(0,2);
a10_raw ~ std_normal(); // normal(0,2);
c11_raw ~ std_normal(); // ~ normal(0,2);
c10_raw ~ std_normal(); // normal(0,2);
b0_raw ~ std_normal(); // ~ normal(0,2);
b1_raw ~ std_normal(); // normal(0,2);
t ~ std_normal();
for (i in 1:N) {
real lps[2];
// logprob conditinal on non-diseased
lps[1] = bernoulli_lpmf(0| p)
+ categorical_lpmf(res1[i]| to_vector(p1[1,1:3,i,1]))
+ categorical_lpmf(res2[i]| to_vector(p1[1,1:3,i,2]))
+ categorical_lpmf(res3[i]| to_vector(p1[1,1:3,i,3])) ;
// logprob conditional on diseased
lps[2] = bernoulli_lpmf(1| p)
+ categorical_lpmf(res1[i]| to_vector(p1[2,1:3,i,1]))
+ categorical_lpmf(res2[i]| to_vector(p1[2,1:3,i,2]))
+ categorical_lpmf(res3[i]| to_vector(p1[2,1:3,i,3])) ;
// marginalize
target += log_sum_exp(lps);
}
}
R code to run model:
res1 <- c(rep(1, 262), rep(2, 24), rep(3, 26))
res2 <- c(rep(1, 183), rep(2, 61), rep(3, 18), rep(1, 13), rep(2, 5),
rep(3, 6), rep(1, 3), rep(2, 4), rep(3, 19))
res3 <- c(rep(1, 181), rep(2, 1), rep(3, 1), rep(1, 56), rep(2, 5),
rep(1,16), rep(2, 1), rep(3, 1), rep(1, 10), rep(3, 3),
rep(1, 5), rep(1, 2), rep(2, 2), rep(3, 2),
rep(1,1), rep(2, 2), rep(3, 4), rep(2, 1), rep(3, 18))
model2 <- stan(file = ......,
data = list(res1=res1, res2=res2, res3=res3, N = 312, nt=3),
iter = 3000,
chains = 1,
control=list(adapt_delta=0.80, max_treedepth =10))