I am modelling data for 28 participants who rated “arousal” and “valence” using a combination pictographic (5 pictures) and Likert-type numeric (9-point) scale for each construct under 7 conditions: auditory only, audio-visual, low-pass filtered auditory only (half of the subjects (n = 14)), low-pass filtered auditory-visual (n = 14) high-pass filtered auditory only (half of the subjects (n = 14)), high-pass filtered auditory-visual (n = 14), and visual only (filtering not applicable). The stimuli consist of 65 “tokens” that have been categorized into “unpleasant”, “neutral”, “pleasant” - the number of tokens in each category is unbalanced.
So, it is a nested design, with half of the participants split on one sub-condition, and a bivariate response
I am trying to model the data as multivariate ordinal (cumulative(logit)).
I have uploaded the data here:
emo_modality_with_all_tokens.csv (609.3 KB)
My code for creating and running the model:
## List required packages ====
Pkgs <- c("tidyverse", "magrittr", "brms",
"rstan", "bayesplot",
"parallel", "tidybayes")
### Load packages ====
lapply(Pkgs, require, c = T)
## Set computing options ====
ncores = detectCores()
rstan_options(auto_write = TRUE)
options(mc.cores = parallel::detectCores())
# Read in data ----
Modality_Ord <- read.csv("emo_modality_with_all_tokens.csv", stringsAsFactors = FALSE) %>%
select("Subject" = ss_id, "Group" = filtering_group, "Filter" = filter_type,
"Condition" = condition, "Category" = category_label,
"Token" = token, "Arousal" = arousal, "Valence" = valence ) %>%
mutate( Category = recode_factor(Category,
unpleasant = "Unpleasant",
neutral = "Neutral",
pleasant = "Pleasant")) %>%
droplevels() %>%
mutate(Condition = ifelse(Condition == "fAO" | Condition == "fAV",
paste(Condition, Filter, sep = "_"), Condition),
Subject = factor(Subject),
Condition = factor(Condition,
levels = c("AO", "fAO_LP", "fAO_HP",
"AV", "fAV_LP", "fAV_HP",
"VO")),
Token = factor(Token),
Arousal = factor(Arousal, ordered = TRUE),
Valence = factor(Valence, ordered = TRUE)) %>%
select(-Group, -Filter) %>%
ungroup() %>%
na.omit()
## Multivariate model formula
Modality_ord_mv_fmla <- bf(mvbind(Arousal, Valence) ~ Category +
Condition +
Category:Condition +
(1|o|Token) +
(1|p|Condition) +
(1|q|Condition:Subject))
## Specify priors
modality_ord_priors_mv <- c(
set_prior("normal(0,2)",
class = "b",
resp = "Arousal"),
set_prior("normal(0,6)",
class = "Intercept",
coef = "",
resp = "Arousal"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Condition",
resp = "Arousal"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Condition:Subject",
resp = "Arousal"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Token",
resp = "Arousal"),
set_prior("normal(0,2)",
class = "b",
resp = "Valence"),
set_prior("normal(0,6)",
class = "Intercept",
coef = "",
resp = "Valence"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Condition",
resp = "Valence"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Condition:Subject",
resp = "Valence"),
set_prior("normal(0,0.125)",
class = "sd",
coef = "Intercept",
group = "Token",
resp = "Valence")
)
# Compile and fit model
Modality_Ord_MV_Mod <-
brm(Modality_ord_mv_fmla,
data = Modality_Ord,
family = cumulative("logit"),
prior = modality_ord_priors_mv,
inits = 0,
iter = 2000,
warmup = 1000,
chains = 2,
cores = ncores,
sample_prior = TRUE,
save_all_pars = TRUE,
control = list(adapt_delta = 0.99,
max_treedepth = 12)
)
The model seems to converge fairly efficiently and the fits to the data for both responses seem good:
Arousal:
pp_check(Modality_Ord_MV_Mod, resp = "Arousal")
Valence:
pp_check(Modality_Ord_MV_Mod, resp = "Valence")
I am seeing some poor PSIS-LOO checks, however:
Looking at Arousal:
loo_mv_arl <- loo(Modality_Ord_MV_Mod, resp = "Arousal",
save_psis = TRUE)
yrep_arl <- posterior_predict(Modality_Ord_MV_Mod, resp = "Arousal")
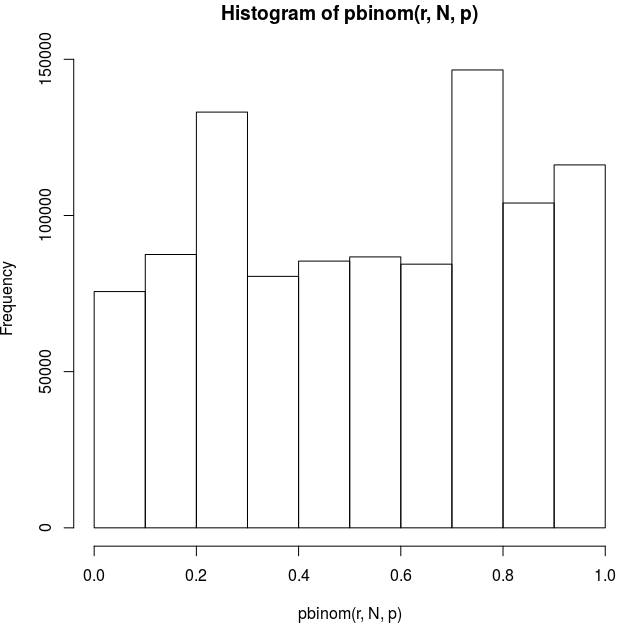
ppc_loo_pit_overlay(
y = as.numeric(Modality_Ord$Arousal),
yrep = as.matrix(yrep_arl),
lw = weights(loo_mv_arl$psis_object)
)
Looking at Valence
loo_mv_val <- loo(Modality_Ord_MV_Mod, resp = "Valence",
save_psis = TRUE)
yrep_val <- posterior_predict(Modality_Ord_MV_Mod, resp = "Valence")
ppc_loo_pit_overlay(
y = as.numeric(Modality_Ord$Valence),
yrep = as.matrix(yrep_val),
lw = weights(loo_mv_val$psis_object)
)
My understanding is that the LOO PITs should be uniform for continuous data; does that apply if the the model is based on a continuous latent variable rather than a true continuous response?
Does this suggest the model is over-dispersed for both responses?
How might I go about further generalizing the model to account for the abundance of large errors?
Would adding a distribution over the disc parameter be of any help?
Edit: I tried adding disc to the model and saw no change in LOO-PIT.