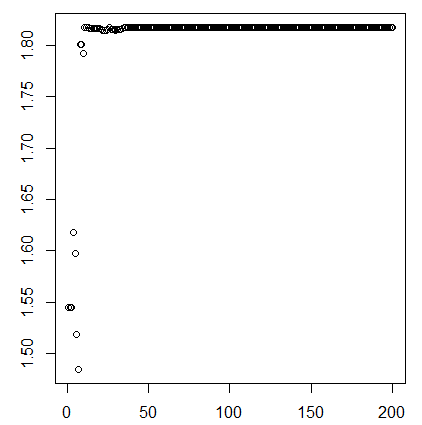
I am encountering divergent transitions when fitting a model involving linear convolution. The actual model is very big, so I am providing a snippet code that highlights that main issue.
First, I create a signal vector as follows.
N = 101
x = -floor(N/2):floor(N/2)
signal = 10*exp(-(10*x/N)^2)
Then, using generated quantities in Stan, I perform a linear convolution of the signal with a Gaussian kernel with a scale parameter and a normal observation process with standard deviation sigma. I use the following inputs
dat <- list(N = N,
signal = signal,
pseudo_pad = 0,
sigma = 1.0,
scale = 2)
to run the following Stan model:
functions{
vector normalize(vector x) {
return x/sum(x);
}
vector Gaussian_kernel(real scale, int M, int buffer){
vector [M] x = linspaced_vector(M, -buffer, buffer);
vector [M] kernel = inv(scale*sqrt(pi())).*exp(-pow(abs(x/scale), 2));
return normalize(kernel);
}
vector conv1D(vector x, vector y){
return abs(inv_fft(fft(x).*fft(y)));
}
vector pad(vector x, int n, int pseudo_pad){
return pseudo_pad == 1 ? append_row(x, zeros_vector(n) + 1e-15): append_row(x, zeros_vector(n));
}
vector linear_convolution(vector signal, vector kernel, int N, int M, int buffer, int pseudo_pad){
vector[N + M - 1] signal_pad = pad(signal, M-1, pseudo_pad);
vector[N + M - 1] kernel_pad = pad(kernel, N-1, pseudo_pad);
vector[N] convolution = segment(conv1D(signal_pad, kernel_pad), buffer+1, N);
return convolution;
}
}
data {
int N;
int pseudo_pad;
vector[N] signal;
real sigma;
real scale;
}
transformed data{
int M = 2 * (N %/% 2) + 1; // this ensures that M is always odd
int buffer = M %/% 2;
}
generated quantities{
vector [N] kernel = Gaussian_kernel(scale, M, buffer);
vector [N] linear_convolution_generated = to_vector(normal_rng(linear_convolution(signal, kernel, N, M, buffer, pseudo_pad), sigma));
}
One thing to note about this code is that I use a variable ‘pseudo_pad’ to decide if I want to add 1e-15 to the zeros_vector(n) when padding signal and kernel vectors. Theoretically, ‘pseudo_pad’ should not have any quantitative effect because compared to the non-padded signal and kernal values 1e-15 is approximately zero.
Then I use the following stan model to recover the parameters scale and sigma:
functions{
vector normalize(vector x) {
return x/sum(x);
}
vector Gaussian_kernel(real scale, int M, int buffer){
vector [M] x = linspaced_vector(M, -buffer, buffer);
vector [M] kernel = inv(scale*sqrt(pi())).*exp(-pow(abs(x/scale), 2));
return normalize(kernel);
}
vector conv1D(vector x, vector y){
return abs(inv_fft(fft(x).*fft(y)));
}
vector pad(vector x, int n, int pseudo_pad){
return pseudo_pad == 1 ? append_row(x, zeros_vector(n) + 1e-15): append_row(x, zeros_vector(n));
}
vector linear_convolution(vector signal, vector kernel, int N, int M, int buffer, int pseudo_pad){
vector[N + M - 1] signal_pad = pad(signal, M-1, pseudo_pad);
vector[N + M - 1] kernel_pad = pad(kernel, N-1, pseudo_pad);
vector[N] convolution = segment(conv1D(signal_pad, kernel_pad), buffer+1, N);
return convolution;
}
}
data {
int N;
int pseudo_pad;
vector[N] signal;
vector[N] convolution_generated;
}
transformed data{
int M = 2 * (N %/% 2) + 1; // this ensures that M is always odd
int buffer = M %/% 2;
}
parameters{
real <lower = 0> scale;
real <lower = 0> sigma;
}
transformed parameters{
vector [N] kernel = Gaussian_kernel(scale, M, buffer);
}
model{
convolution_generated ~ normal(linear_convolution(signal, kernel, N, M, buffer, pseudo_pad), sigma);
[sigma, scale] ~ exponential(1);
}
with the following input values:
dat <- list(N = N,
signal = signal,
pseudo_pad = pseudo_pad,
convolution_generated = convolution_generated)
The convolution_generated is the output of the generated quantities.
The problem is when I use pseudo_pad = 0, I get divergent transitions:
All 4 chains finished successfully.
Mean chain execution time: 124.1 seconds.
Total execution time: 163.7 seconds.
Warning: 909 of 1600 (57.0%) transitions ended with a divergence.
See https://mc-stan.org/misc/warnings for details.
Warning: 691 of 1600 (43.0%) transitions hit the maximum treedepth limit of 10.
See https://mc-stan.org/misc/warnings for details.
# A tibble: 2 × 10
variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 scale 1.37 1.15 0.897 0.879 0.434 2.72 Inf 4.08 NA
2 sigma 1.02 1.12 0.285 0.133 0.543 1.30 Inf 4.08 NA
But when I use pseudo_pad = 1, the fitting goes well.
All 4 chains finished successfully.
Mean chain execution time: 3.2 seconds.
Total execution time: 3.7 seconds.
# A tibble: 2 × 10
variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 scale 2.20 2.20 1.45 1.83 0.102 4.59 1.01 291. 439.
2 sigma 1.02 1.02 0.0709 0.0678 0.912 1.14 1.01 393. 386.
I was wondering if someone could explain why zero padding is leading to divergent transitions.
Thank you in advance