Hello,
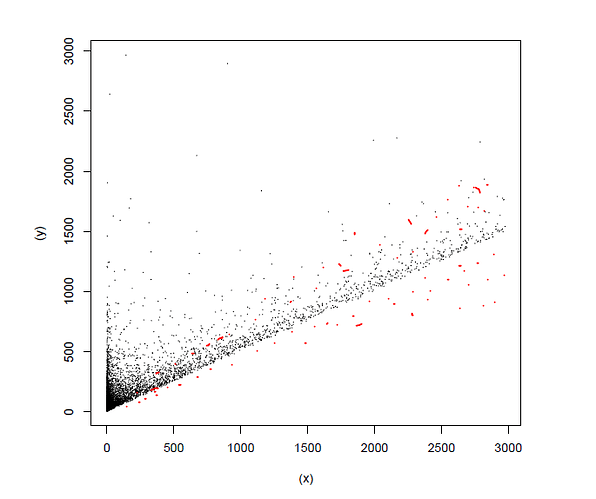
I have a system where the noiseless data (BLACK dots) has a half normal-like regression
While the noise (RED dots) is heteroshedastic, and breaks the truncated data structure (or not?)
- Should I use a truncated linear regression?
- And what about the heterochedastic noise?
- In case, what is the best way to code truncate linear regression (
y[n] ~ normal(x[n] * a, sigma) T[??,];)
Thanks
There’s a lot going on near small values, have you looked at the log-log plot for this data? It’s hard to say much without visualizing this more.
Yes,
even if the density is higher for small values it’s still linear.
At the moment I am using log log regression (especially for heterochedasticity), however for some low values there is some complication
Here is the log-log plot (noiseless)
I’m pretty sure you’ve posted this model before (or somebody else is working with the same data (?)). Doing a truncated model might work well if you have an expression for that line, it’s clearly not just half-normal on the lower end.
Quite some time ago, and it’s working well, in log-log version, but now before to close test I want to do the last check if better model is possible. (I approached truncated models just recently, and I had this thing in mind)
Do you mean log plot or raw plot? The log plot the line follows log(b0 + x * b1)
I would like truncated on raw data (NON-log transformed). What do you think?
I think the joy of discovery is well worth it.
Eheh, Yes at this stage the joy of discovery should be replaced by the joy of publishing. But I will take some time to rule this possibility in/out.
The question to start from something is:
can in principle, in this format y[n] ~ normal(x[n] * a, sigma) T[??,];
?? be replaced with regression variables T[x[n] * a,] ?
Thanks
Think about it this way:
target += normal_lpdf(y[n] | x[n] * a, sigma)
+ normal_cdf(x[n] * a | x[n] * a, sigma);
Sampler provides a and sigma, you provide x and y. It’s a density and life goes on. I don’t think this holds as a good model for the lower end of your tail so I doubt it will work but I always have to plot more.