I tried a couple models, but there was enough variation in the stepsizes I didn’t really know what to think. I can run some more in a few days and try to get a handle on the noise, if that’s helpful. Were we just expecting the stepsizes to be larger?
Thanks!
The step size should increase uniformly, kind of by construction in the modified code. The one circumstance where you might not see the step size increase is if the adaptation is noisy and you see accept_stat higher with the new code than with the nominal code (i.e. for the same accept_stat the new code should always give a higher step size than the nominal code).
This should lead to fewer leapfrog steps needed to simulate the optimal integration times in each trajectory, and hence cheaper transitions overall, but I’m seeing longer trajectories in some circumstances which may be a consequence of the generalized No-U-Turn criterion missing a u-turn due to the less accurate numerical trajectories.
Not only data set, but also the function of the parameters being evaluated.
The code’s there, which eliminates all ambiguity, but it’s hard to read.
I’d also like a rough description of what’s changing.
The place to start for the math is @betanalpha’s paper on optimal integration time for HMC. That’ll cite the Beskos et al. paper on tuning HMC. The other thing to read is the newer paper by Livingstone et al. on kinetic energy choice. These issues are all related.
P.S. What I’d like to do to fix adaptation, since we’re on the topic, is make it more continuous with old draws being discounted rather than thrown away in blocks.
Incorrect. It cites Optimizing The Integrator Step Size for Hamiltonian Monte Carlo which supersedes the Beskos et al paper with a more general result and a critical upper bound.
I played around with this but in early adaptation the sampler moves fast enough (and we don’t have enough iterations) that the old samples are hard to down weight sufficiently strongly. The windows were introduced to let the sampler warm up in a small part of the typical set without burning the properties of that neighborhood into the variance estimators permanently.
Anyone running anything or planning to run anything? I have a suite of tests demonstrating the relevant behaviors, including those manifested in @jroon’s results, but the more comparisons the better.
For example this is the relevant summary info,
100 Dimensional IID Normal
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.3e-01 2.0e-03 1.5e-01 0.56 8.6e-01 1.0
stepsize__ 4.8e-01 1.3e-14 9.4e-15 0.48 4.8e-01 0.48
treedepth__ 3.0e+00 nan 5.1e-14 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 nan 9.7e-14 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+02 2.5e-01 1.0e+01 83 1.0e+02 117
697300 total effective samples
19.922 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.4e-01 1.8e-03 1.3e-01 0.60 8.7e-01 1.0
stepsize__ 5.4e-01 5.3e-15 3.8e-15 0.54 5.4e-01 0.54
treedepth__ 3.0e+00 nan 5.1e-14 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 nan 9.7e-14 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+02 2.4e-01 9.9e+00 84 1.0e+02 117
932700 total effective samples (33% improvement)
26.648 effective samples per gradient evaluation (34% improvement)
Sat down this afternoon to get the results of my tests parsed. I’ll get something up soon.
Of course it’s entirely possible I screwed up the benchmarks and need to run them again but we’ll see :/.
Alright here goes. It’s the 6 models from the adaptation paper (https://arxiv.org/abs/1905.11916). 32 runs each. I ran them with dense and diagonal adaptation with and without the new stepsize adaptation.
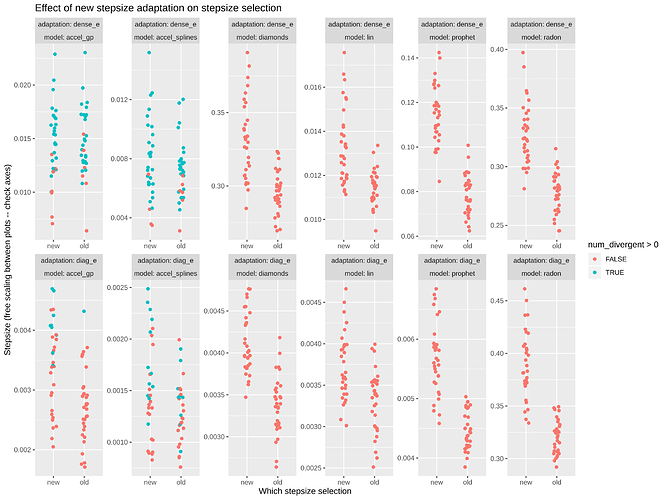
Here’s a plot of the effect on the stepsize (edit: watch the scales and the zeroes in these plots):
Here’s a plot of the effect on the effective sample size/s (these are sketchy and I should recompute them in groups – it’s easy for the single chain ESS/s to be screwy, edit: watch the scales and the zeroes in these plots):
Here’s a csv with the datapoints if you want to plot them differently:
stepsizes_df.csv (42.4 KB)
Here’s code to reproduce the plots:
library(ggplot2)
df = read_csv("stepsizes_df.csv")
df %>%
ggplot(aes(which_stepsize, stepsize)) +
geom_jitter(aes(color = num_divergent > 0), width = 0.1) +
facet_wrap(adaptation ~ model, nrow = 2, scales = "free", labeller = label_both) +
xlab("Which stepsize selection") +
ylab("Stepsize (free scaling between plots -- check axes)") +
ggtitle("Effect of new stepsize adaptation on stepsize selection")
ggsave("stepsize.png", width = 12, height = 9, dpi = 600)
df %>%
ggplot(aes(which_stepsize, min_neffps)) +
geom_jitter(aes(color = num_divergent > 0), width = 0.1) +
facet_wrap(adaptation ~ model, nrow = 2, scales = "free", labeller = label_both) +
xlab("Which stepsize selection") +
ylab("Minimum ESS/s (free scaling between plots -- check axes)") +
ggtitle("Effect of new stepsize adaptation on min ESS/s\n(ESS/s calculations are for single chains)")
ggsave("effective_sample_size.png", width = 12, height = 9, dpi = 600)
The only thing that really alarms me are the radon calculations that have very low ESS/s values. Otherwise it looks like the stepsize is being pushed larger without any downside. This translates to much higher ESS/s in a few cases. Outside of the Radon example, doesn’t seem like this makes things worse.
It looks like the radon models were bumping up against treedepth things? I dunno what was happening. I assume it’s something blowing up early in adaptation, but I didn’t check.
The model is (from here: https://mc-stan.org/users/documentation/case-studies/radon.html): radon.stan (371 Bytes)
With data: radon.data.R (585.3 KB)
I was able to reproduce the problem with:
radon sample data file=radon.data.R init=2.0 random seed=3669238606
Here’s the output csv for that run: output.csv (3.2 MB)
Alright now, there are benchmarks – what’s the new adaptation doing differently?
Great, thanks, @bbbales2! This is pretty much consistent with my expectations. Note that to avoid the ESS/time variability it’s often easier to compare ESS/n_leapfrog – because the autodiff is fixed the only algorithmic differences will be in the number of leapfrog/gradient calls and not the time per each of those calls.
Additionally I avoid the older examples like the BUGS models and the radon model as the priors and parameterizations are terrible. Similarly I don’t consider anything model that yields divergences in performance comparisons – once it’s divergent the performance is irrelevant.
When building up empirical comparisons I instead focus on models that capture single qualitative features of interest – high dimension, high linear correlations, heavy tails, etc – as they make it easier to identify which features, if any, might cause problems.
For this tweak my comparison were
10 Dimensional IID Normal
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.9e-01 1.5e-03 1.1e-01 0.67 9.2e-01 1.0
stepsize__ 6.8e-01 1.7e-14 1.2e-14 0.68 6.8e-01 0.68
treedepth__ 2.8e+00 5.6e-03 3.7e-01 2.0 3.0e+00 3.0
n_leapfrog__ 6.6e+00 1.8e-02 1.2e+00 3.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+01 7.2e-02 3.2e+00 5.4 9.7e+00 16
99700 total effective samples
2.848 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.7e-01 1.6e-03 1.3e-01 0.61 9.1e-01 1.0
stepsize__ 8.2e-01 5.7e-15 4.0e-15 0.82 8.2e-01 0.82
treedepth__ 3.9e+00 2.6e-02 1.7e+00 2.0 4.0e+00 6.0
n_leapfrog__ 3.5e+01 6.4e-01 4.0e+01 3.0 2.3e+01 127
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+01 6.8e-02 3.2e+00 5.5 9.7e+00 16
51700 total effective samples (48% reduction)
1.477 effective samples per gradient evaluation (48% reduction)
100 Dimensional IID Normal
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.3e-01 2.0e-03 1.5e-01 0.56 8.6e-01 1.0
stepsize__ 4.8e-01 1.3e-14 9.4e-15 0.48 4.8e-01 0.48
treedepth__ 3.0e+00 nan 5.1e-14 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 nan 9.7e-14 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+02 2.5e-01 1.0e+01 83 1.0e+02 117
697300 total effective samples
19.922 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.4e-01 1.8e-03 1.3e-01 0.60 8.7e-01 1.0
stepsize__ 5.4e-01 5.3e-15 3.8e-15 0.54 5.4e-01 0.54
treedepth__ 3.0e+00 nan 5.1e-14 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 nan 9.7e-14 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.0e+02 2.4e-01 9.9e+00 84 1.0e+02 117
932700 total effective samples (33% improvement)
26.648 effective samples per gradient evaluation (34% improvement)
100 Dimensional IID Student t (5 degrees of freedom)
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.8e-01 1.5e-03 1.1e-01 0.67 9.1e-01 1.00
stepsize__ 3.3e-01 9.4e-16 6.7e-16 0.33 3.3e-01 0.33
treedepth__ 4.0e+00 2.0e-04 1.4e-02 4.0 4.0e+00 4.0
n_leapfrog__ 1.5e+01 1.7e-02 1.1e+00 15 1.5e+01 15
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.2e+02 3.2e-01 1.2e+01 98 1.2e+02 136
812700 total effective samples
23.220 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.8e-01 1.7e-03 1.2e-01 0.65 9.2e-01 1.00
stepsize__ 3.6e-01 9.6e-15 6.8e-15 0.36 3.6e-01 0.36
treedepth__ 4.1e+00 6.0e-03 3.8e-01 4.0 4.0e+00 5.0
n_leapfrog__ 2.0e+01 2.5e-01 1.5e+01 15 1.5e+01 47
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 1.2e+02 3.2e-01 1.2e+01 97 1.2e+02 135
815300 total effective samples (0.3% improvement)
23.294 effective samples per gradient evaluation (0.3% improvement)
50 Dimensional Multi Normal, rho=0.25
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.4e-01 1.9e-03 1.4e-01 0.57 8.7e-01 1.0
stepsize__ 5.1e-01 2.8e-15 2.0e-15 0.51 5.1e-01 0.51
treedepth__ 3.0e+00 2.0e-04 1.4e-02 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 3.6e-03 2.5e-01 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 5.0e+01 1.7e-01 7.2e+00 39 5.0e+01 62
364900 total effective samples
10.425 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.5e-01 1.8e-03 1.3e-01 0.60 8.9e-01 1.0
stepsize__ 5.5e-01 6.9e-15 4.9e-15 0.55 5.5e-01 0.55
treedepth__ 3.0e+00 nan 5.1e-14 3.0 3.0e+00 3.0
n_leapfrog__ 7.0e+00 nan 9.7e-14 7.0 7.0e+00 7.0
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 5.0e+01 1.5e-01 7.1e+00 39 5.0e+01 62
437100 total effective samples (20% improvement)
12.488 effective samples per gradient evaluation (20% improvement)
10 Dimensional Multi Normal, rho=0.80
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 9.0e-01 1.2e-03 9.8e-02 0.70 9.3e-01 1.0
stepsize__ 3.1e-01 8.6e-15 6.1e-15 0.31 3.1e-01 0.31
treedepth__ 3.5e+00 1.0e-02 6.9e-01 3.0 4.0e+00 5.0
n_leapfrog__ 1.7e+01 1.4e-01 9.7e+00 7.0 1.5e+01 31
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 9.9e+00 7.3e-02 3.1e+00 5.4 9.6e+00 15
20100 total effective samples
0.574 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 9.1e-01 1.1e-03 9.0e-02 0.73 9.4e-01 1.0
stepsize__ 2.9e-01 7.9e-17 5.6e-17 0.29 2.9e-01 0.29
treedepth__ 3.5e+00 1.1e-02 7.0e-01 3.0 3.0e+00 5.0
n_leapfrog__ 1.7e+01 1.5e-01 9.9e+00 7.0 1.5e+01 31
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 9.9e+00 7.4e-02 3.1e+00 5.3 9.6e+00 15
19200 total effective samples (4% reduction)
0.548 effective samples per gradient evaluation (4% reduction)
50 Dimensional Multi Normal, rho=0.80
Nominal:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.6e-01 1.6e-03 1.2e-01 0.63 8.9e-01 1.00
stepsize__ 2.3e-01 5.4e-15 3.8e-15 0.23 2.3e-01 0.23
treedepth__ 4.7e+00 1.3e-02 8.0e-01 3.0 5.0e+00 6.0
n_leapfrog__ 4.0e+01 3.7e-01 2.4e+01 15 3.1e+01 79
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 5.0e+01 1.7e-01 7.0e+00 39 5.0e+01 62
182400 total effective samples
5.211 effective samples per gradient evaluation
Adapt Tweak:
Mean MCSE StdDev 5% 50% 95%
accept_stat__ 8.4e-01 1.9e-03 1.4e-01 0.56 8.7e-01 1.00
stepsize__ 2.8e-01 3.1e-16 2.2e-16 0.28 2.8e-01 0.28
treedepth__ 4.7e+00 1.1e-02 7.0e-01 4.0 5.0e+00 6.0
n_leapfrog__ 3.7e+01 3.9e-01 2.5e+01 15 3.1e+01 79
divergent__ 0.0e+00 nan 0.0e+00 0.00 0.0e+00 0.00
energy__ 5.0e+01 1.6e-01 6.9e+00 39 5.0e+01 62
229000 total effective samples (26% improvement)
6.542 effective samples per gradient evaluation (26% improvement)
Hopefully the pattern is clear and consistent with what you see. In high-dimensions the tweak improves performance unless the tails are really heavy, in which case the performance is the same. In low dimensions the tweak can reduce performance because the tree depth distribution expands and the sampler wastes time on overly long trajectories. This is due to the higher step size leading to the generalized No-U-Turn criterion to be evaluated at too crude a resolution, with the numerical trajectories continuing much too far before finally being terminated. At the same time this performance dip is in low dimensional models which already run extremely quickly.
The requisite reading for what’s going on is https://arxiv.org/abs/1411.6669. There we showed that in the original Hybrid Monte Carlo scheme, where the numerical Hamiltonian trajectory is used as a Metropolis proposal, the optimal step size is the one that yields an average Metropolis acceptance probability of 0.8.
Both NUTS and the current implementation of Hamiltonian Monte Carlo used in Stan, however, are not Hybrid Monte Carlo schemes. Instead they sample from an entire trajectory whose length is grown dynamically around the initial point. Because of that the above analysis isn’t immediately applicable.
In order to use the above step size optimization criterion analysis one has to come up with a proxy Metropolis acceptance probability (incidentally this is which is why it’s called “accept_stat” and not “accept_prob” in the diagnostics). The current version of Stan pretends that each point in a given numerical trajectory could have been a Metropolis proposal and averages over the corresponding Metropolis acceptance probabilities uniformly.
We do not, however, sample states within a numerical trajectory uniformly in the algorithm. Instead we sample them based on their weights, e^{H(q, p)}. By weighting the fake Metropolis acceptance probabilities uniformly we over emphasize those states that are unlikely to be chosen, and end up with a conservative proxy acceptance statistic.
This new branch considers each state as a proxy proposal but weights the corresponding acceptance probabilities by the state weight, e^{H(q, p)}. In this way those states that are likely to be chosen have a greater influence on the proxy acceptance statistic, and any state with vanishing weight is not considered at all. This will always lead to a higher proxy acceptance stat for a given step size, which will cause the adaptation to push toward higher step sizes and, for a fixed integration time, cheaper numerical trajectories.
The effect of this change should be most evident in higher dimensions where the change in Hamiltonian values along a numerical trajectory increase. At the same time the optimality analysis is most accurate in higher dimensions (in the paper we utilize an expansion in step sizes but the coefficients of that expansion implicitly depend on dimension). This is the same as the 0.234 analysis of Roberts et al which is a purely asymptotic result and isn’t optimal for lower dimensional targets.
That said, the real issue here is that forcing the step size to larger values makes the generalized No-U-Turn criterion less accurate. Combined with the multiplicative trajectory expansion this can result in a heavy tail of longer numerical trajectories, especially in lower dimensions. Once again this is confirmed in the empirical results.
Ultimately the theory, which is better posed in higher dimensions, strongly motivates the reweighed acceptance statistic, which is backed up by empirical results. The cost is lower performance of low dimensional models, but these models run so quickly that the added cost isn’t particularly relevant.
Tomorrow I’ll update all of the tests and submit an issue/PR. Thanks again for the help with the empirical tests @jroon and @bbbales2!
Oh yeah, that would be a better way to do it.
I’m looking at this, by the way. The plan was to multiplicatively expand up to some treedepth, and then additively expand with trees of that treedepth (until some maximum number of leapfrog steps). Aki said you’d mentioned something like this?
Fair, but this model did work pretty well with the last sampler.
I had a look at the outputs of one chain that ran fast and one didn’t. The timestep adaptation doesn’t look like it’s going crazy (not crashing to zero or anything). The outputs of the two chains mix fine (according to Rhat), and the mass matrices are very close (all the individual entries are within a factor of two of each other).
I think this is worth investigating.
The divergence plots are there to show that this didn’t appear to be increasing the number of them.
This would handle them differently, right? Trajectories marked divergent are ones where there are leapfrog steps with huge energy errors? In the original method, these points would have close to a zero Metropolis accept probability and would cause the timestep to get smaller.
And what’d happen with this method? The leapfrog steps with huge energy error would just influence the timestep less? This makes me think if there was potential for divergences before, it’d be a little bit higher now. These tests weren’t really enough to probe this I guess.
The problem is that waiting to additively expand until later doesn’t offer much computational advantage. You still have to save every state in the trajectory in order to accept those later additive expansions, and you still have to check the new state against every existing state. Doing a multiplicative expansion first just saves some relatively inexpensive calculations, not the really costly ones.
Yeah, really all of the old BUGS models need investigating!
If the inverse metrics and step sizes are nearly the same and there aren’t divergences then I don’t know what would be causing huge time differences.
Again the biggest issue with touching the step size concerns the dynamic trajectory expansion. Firstly by decreasing the step size you can hit a threshold where you need a little bit more integration time but can only achieve that by performing an entire trajectory doubling (the max cost difference here is then only about 2). By increasing the step size you can then potentially get significant speed ups if you were just past the threshold before, but at the same time larger step sizes make the termination criterion less accurate and hence you might accidentally expand past the point you care about. But again the speed difference should be less than a factor of 2.
Right – the divergent end of the trajectory would be downweighted in computing the accept stat, leading to higher accept stats and the adaptation pushing to higher step sizes. But the trajectory would still be marked as divergent, which is the most important thing.
The resolution, however, would still be the same. If the neighborhood of high curvature is sufficiently small then you might be able to get away with a higher adapt_delta but in most cases you’ll need to improve your priors and parameterizations. Controversial stance – if you need to run with adapt_delta > 0.9 then your model isn’t as identified as you want it to be.
Ultimately the number of divergences isn’t as important as whenever you see divergences or not.
Oh, well, I still gotta learn how all this treebuilding stuff works I s’pose. Otherwise it’ll always be handwavy to me.
Same. I thought it would be obvious once I popped open the save_warmup=1 csv, but it wasn’t.
The code changes are going to be simple enough to review, but can you have a look into this before I get to the Github review? Since this is an algorithm change we’ll have to talk to everyone before we incorporate it. Best that the case be as clear as possible.
If this is an unrelated, pre-existing issue that’s just been uncovered by setting the stepsize higher (which seems like what it is), that’s cool, we can just document it and set it aside. But I’d like to know how to trigger it.
I’ll run some tests in a couple days. I suspect we can trigger this just by using fixed adaptations and larger and larger stepsizes.
Whoops, realized I already had the data for this. Check out this plot (edit: these are all numbers for the radon model):
Time to run some more tests!
I’m about to board a flight to London and will respond proper tomorrow but if you plot all the combinatorics of {ess, ess per time, n_leapfrog, stepsize} then the underlying problem should be pretty evident.
Yup got it reproduced in a regular cmdstan and rstan by just lowering the adapt_delta so the timestep goes higher and also reproduced it outside of Stan. So this is definitely a problem that is outside the scope of this pull.
Hi Michael,
I tried out the test on a slightly more complicated example but in this model it does not seem to make a lot of change. Below the results and the Julia program I used.
RequestMichael.txt (8.4 KB)
lba.jl.txt (8.6 KB)
Rob
Thanks. While the overall affect is small the step size is behaving as expected. Perhaps the biggest issue in this particular example is that the adaptation for the step size isn’t really reaching a good equilibrium in either case – the average acceptance stat is much bigger than the expected 0.8. Unless you have a non-default configuration of the sampler?
I tried to document everything with both text and figures in the conceptual paper but I’m happy to sit down and walk you through it when you get to town.
Definitely the default configuration:
julia> stanmodel.cmds[1]
/var/folders/fb/m164mwfn3v1btnrpmmyzpvqh0000gn/T/jl_tZ63kH/LBA sample num_samples=1000 num_warmup=1000 save_warmup=0 thin=1 adapt engaged=1 gamma=0.05 delta=0.8 kappa=0.75 t0=10.0 init_buffer=75 term_buffer=50 window=25 algorithm=hmc engine=nuts max_depth=10 metric=diag_e stepsize=1.0 stepsize_jitter=1.0 random seed=-1 init=2 id=1 data file=/var/folders/fb/m164mwfn3v1btnrpmmyzpvqh0000gn/T/jl_tZ63kH/LBA_data_1.R output file=/var/folders/fb/m164mwfn3v1btnrpmmyzpvqh0000gn/T/jl_tZ63kH/LBA_chain_1.csv refresh=100
Interesting, thanks for the hint.
Odd that the step size adaptation isn’t equilibrating well – I wonder if it’s the truncation in the normal priors inducing any nasty geometry. In any case for a model with so few parameters having no effect is actually a pretty good success!