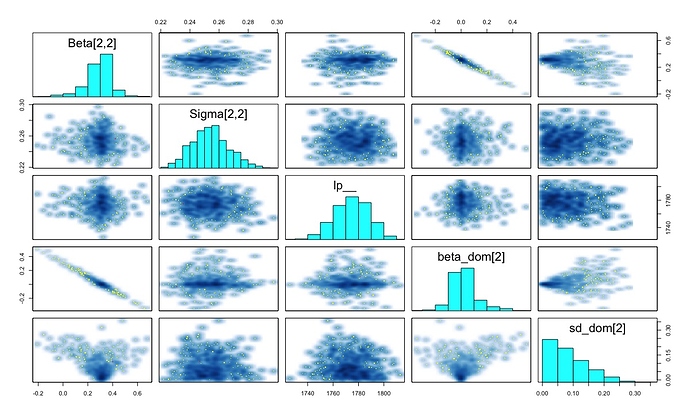
Hi all. So I’ve been working on the model below for a few days now as an attempt to implement the models described here - nested domain multivariate hierarchical models. With simulated data the models are more or less recovering the correct answers, however in the current form I’m getting max treedepth warnings on virtually every iteration. On increasing the max-treedepth to 12. I’m still getting these errors 950 iterations out of 1000, although traceplots look a bit better. Its also super-slow. I’ve tried a few different things including re-parametrisation and altering the priors, but its not helping.
Stuff I tried that didnt’ help is commented out in the code below. If I remove the beta_oc, beta0_dom and beta_dom terms the model behaves very well, but then I have no nesting of outcomes in domains which is what I want! However that makes me think the problem lies with those terms, although I’ve been unable to figure it out.
Model code below and I include a file for simulating data.
Any suggestions folks? Thanks in advance!
Sim data: gen_data_y3_x3.R (1011 Bytes)
Stan model:
data{
int<lower=0> NvarsX; // num independent variables
int<lower=0> NvarsY; // num dependent variables
int<lower=0> N; // Total number of rows
int<lower=0> N_ind; // Number of individuals/participants
int<lower=2> N_dom; // Number of domains in which outcomes are nested
int indiv[N]; // data to identify individuals
matrix[N, NvarsX] x; // data for independent vars
vector[NvarsY] y [N]; // data for dependent vars
int dom[NvarsY]; // data to define outcome domains
}
parameters{
cholesky_factor_corr[NvarsY] L_Omega; //For the correlations between outcome variables
vector<lower=0>[NvarsY] L_sigma; //Residual SD of outcome variables
matrix[NvarsY, N_ind] Zbeta0_ind; //Intercepts at individual level
matrix[NvarsX, N_ind] Zbeta_ind [NvarsY]; //Coefficients at the individual level
vector<lower=0> [NvarsY] Tau0; //Random effect for individual intercepts
matrix<lower=0> [NvarsY, NvarsX] Tau; //Random effect for indvidiual coefficients
vector[NvarsY] Beta0; // Level 2 intercepts for each Y
vector[NvarsX] Beta [NvarsY]; // Level 2 coefficients for each X vs each Y
vector[NvarsY] beta_oc; // Outcome specific random effect
//vector[NvarsY] zb_oc;
//vector[NvarsY] Tau_oc;
vector[N_dom] beta0_dom; // Domain specificrandom intercept
vector[N_dom] beta_dom; // Domain specific random effect
vector<lower = 0> [NvarsY] sd_oc; // SD for outcome effect
vector<lower = 0> [N_dom] sd_dom; // SD for domain effect
vector<lower = 0> [N_dom] sd0_dom; // SD for domain intercept
}
transformed parameters{
vector[NvarsY] mu[N];
matrix[NvarsY, NvarsY] L_Sigma;
matrix[NvarsY, N_ind] beta0_ind;
matrix[NvarsX, N_ind] beta_ind [NvarsY];
L_Sigma = diag_pre_multiply(L_sigma, L_Omega);
// Define Individual level betas
for (s in 1:N_ind){
for (dv in 1:NvarsY){
// Non centered specification for component in ( )
beta0_ind[dv, s] = ( Zbeta0_ind[dv, s] * Tau0[dv] + Beta0[dv] ) +
beta0_dom[ dom[dv] ];
for (iv in 1:NvarsX){
// Non centered specification for component in ( )
beta_ind[dv, iv, s] = ( Zbeta_ind[dv, iv, s] * Tau[dv, iv] + Beta[dv, iv] ) +
beta_oc[dv] + beta_dom[ dom[dv] ];
//(zb_oc[dv] * Tau_oc[dv] + beta_oc[dv] ) + beta_dom[ dom[dv] ];
}
}
}
// Define level 2 betas
for (i in 1:N){
for (dv in 1:NvarsY){
mu[i, dv] = beta0_ind[dv, indiv[i]] +
dot_product( to_vector( beta_ind[dv, 1:NvarsX, indiv[i]] ), x[i, 1:NvarsX] );
}
}
}
model{
//Priors
L_Omega ~ lkj_corr_cholesky(1);
L_sigma ~ cauchy(0, 5);
Tau0 ~ cauchy(0, 2);
to_vector(Tau) ~ cauchy(0,2);
to_vector(beta_oc) ~ normal(0, sd_oc);
//to_vector(zb_oc) ~ normal(0, 1);
//to_vector(Tau_oc) ~ normal(0, 2);
to_vector(beta_dom) ~ normal(0, sd_dom);
to_vector(beta0_dom) ~ normal(0, sd0_dom);
to_vector(sd_oc) ~ normal(0, 0.5);
to_vector(sd_dom) ~ normal(0, 0.5);
to_vector(sd0_dom) ~ normal(0, 0.5);
for (s in 1:N_ind){
for (dv in 1:NvarsY){
Zbeta0_ind[dv, s] ~ normal(0, 1);
Zbeta_ind[dv, 1:NvarsX, s] ~ normal(0, 1);
}
}
to_vector(Beta0) ~ cauchy(0, 5);
//for( dv in 1:NvarsY){
// to_vector(Beta[dv, 1:NvarsX]) ~ cauchy(0, 2);
//}
//likelihood
for (i in 1:N){
y[i, 1:NvarsY] ~ multi_normal_cholesky(mu[i, 1:NvarsY], L_Sigma);
}
}
generated quantities{
matrix[NvarsY, NvarsY] Omega;
matrix[NvarsY, NvarsY] Sigma;
Omega = multiply_lower_tri_self_transpose(L_Omega);
Sigma = quad_form_diag(L_Omega, L_sigma);
}