Hi,
There seems to be an issue with ordinal models whereby comparing two groups with identical data leads to non-zero estimates of effects on the latent mean and standard deviation.
Here’s a reproducible example using these two packages
library(tidyverse)
library(brms)
Here’s some example data, as you can see they are identical across groups
dat <- tribble(
~ID, ~Rating, ~Count,
12, 1, 1,
12, 2, 1,
12, 3, 2,
12, 4, 6,
12, 5, 22,
5, 1, 1,
5, 2, 1,
5, 3, 2,
5, 4, 6,
5, 5, 22
) %>%
mutate(ID = factor(ID))
I then estimate a pretty standard cumulative model where I allow the mean and variance of the latent variable to be different for the two groups. The only nonstandard thing here is using weights (so if I have massive data the model still runs fast) but it makes no difference here.
fit <- brm(
bf(Rating | weights(Count) ~ 1 + ID) + lf(disc ~ 0 + ID , cmc=FALSE) ,
family = cumulative("probit") ,
data = dat,
)
summary(fit, prior = TRUE)
I would obviously expect effects on the mean and SD to be zero because the data are identical, but here are the results:
Family: cumulative
Links: mu = probit; disc = log
Formula: Rating | weights(Count) ~ 1 + ID
disc ~ 0 + ID
Data: dat (Number of observations: 10)
Samples: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup samples = 4000
Priors:
Intercept ~ student_t(3, 0, 2.5)
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept[1] -2.12 0.43 -3.09 -1.36 1.00 1866 2272
Intercept[2] -1.64 0.33 -2.33 -1.02 1.00 2754 2940
Intercept[3] -1.17 0.26 -1.71 -0.68 1.00 4355 2995
Intercept[4] -0.43 0.23 -0.88 0.02 1.00 3939 3618
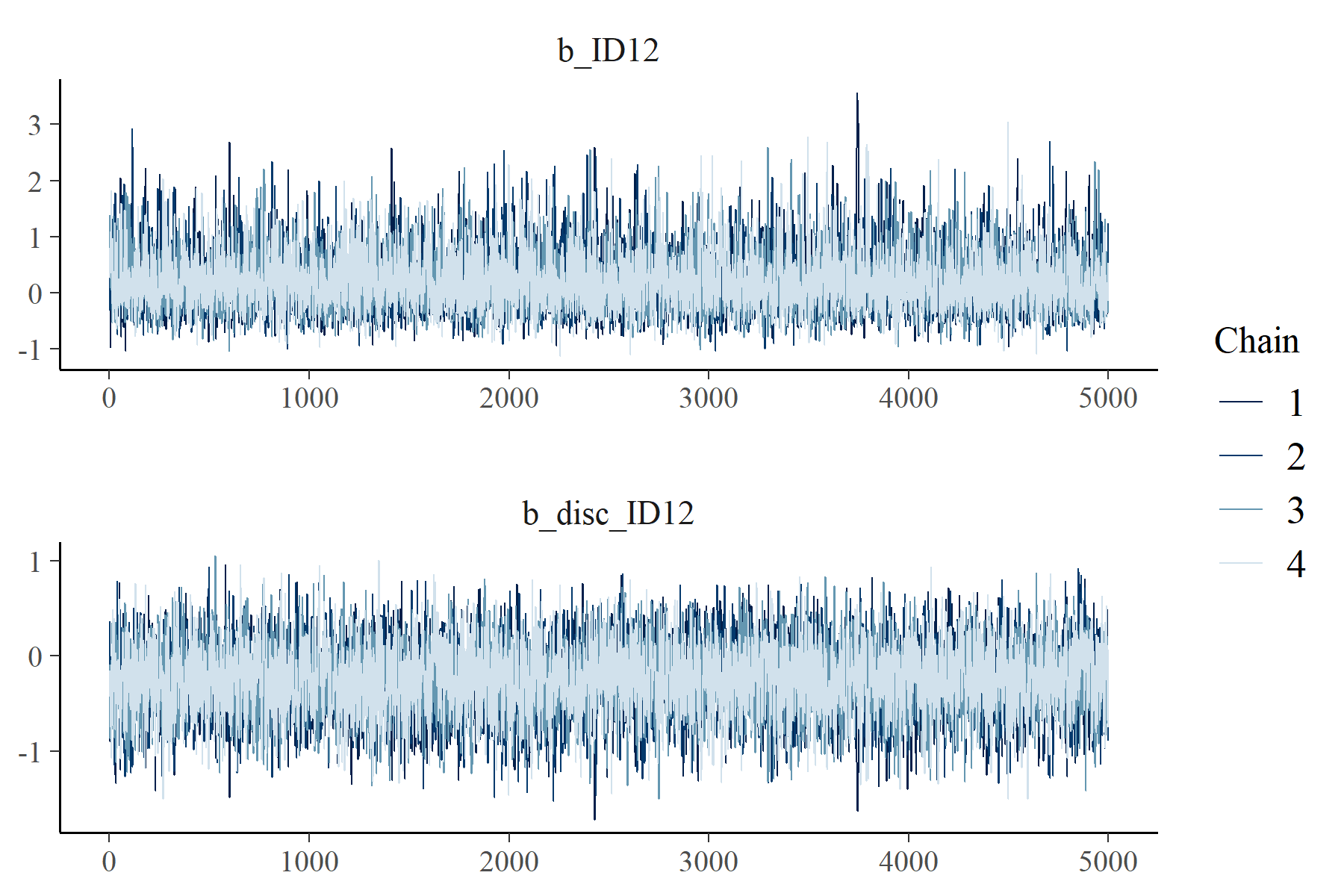
ID12 0.25 0.53 -0.57 1.52 1.00 1426 1598
disc_ID12 -0.25 0.37 -0.98 0.47 1.00 1192 1970
Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).
This is pretty surprising behaviour (to me at least). I talked about it with @paul.buerkner (who probably has a thing or two to say about it), but if anyone else has insight into why this is happening, I’m all ears. Thanks!