I am running an analysis based closely on this notebook by Aki Vehtari. I want to estimate the effect of amphetamine use prior to start of treatment on number of days of cannabis use during treatment among people enrolled in an opioid treatment program.
The data come from routinely collected outcomes data from clients enrolled in opioid agonist treatment at public drug and alcohol clinics. The goal is to examine the effects of amphetamine use prior to starting treatment on clinical outcomes during treatment in this cohort. The outcome variables in this model is outcome and is a bounded count indicating days of cannabis use in the previous 28 day period. The predictors are ats_baseline indicating days used in the 28 days prior to starting treatment and yearsFromStart indicating time in years that the outcome variable was observed. Using a betabinomial model the goal is to estimate differences in days of cannabis use at various points throughout the first year of treatment based on candidate values of ats_baseline at baseline and then three-monthly intervals during the first year of treatment. yearsFromStart and the interaction between yearsFromStart and ats_baseline are represented by a spline. set is a variable indicating the number of trials for each measure (always 28).
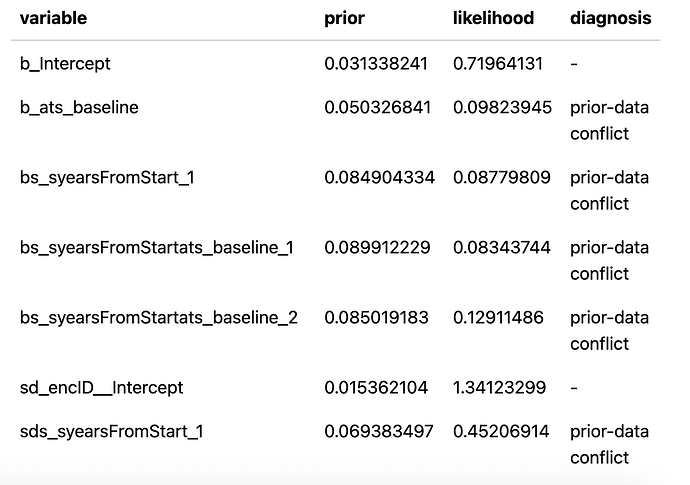
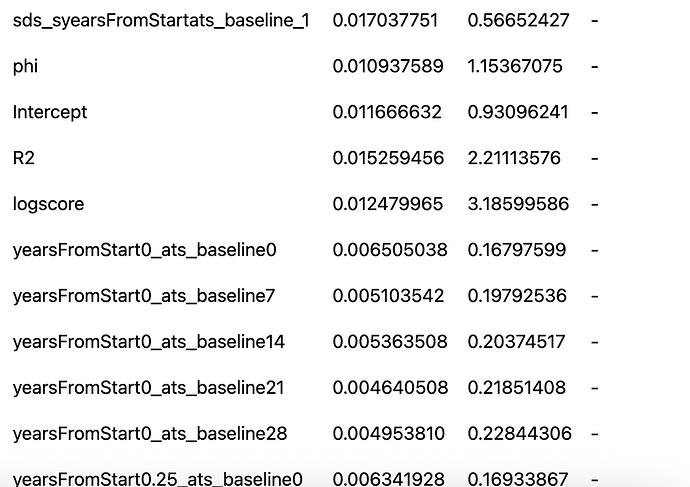
When I originally ran this model I used the default priors in brms. The model converged ok but when I did a prior sensitivity check using the priorsense package I got several prior-data conflicts. Here is the output
Professor Vehtari advised than in the case of a spline model is was better to perform the prior-scaling sensitivity analysis on the predictions rather than the posterior estimates, and referred me to this vignette detailing how to power-scale priors in a model with splines. This involves specifying priors on betas and the sds parameters for the splines and inclusion of the tag = argument in each of the prior specifications.
Here is the data.
amphUseInOTP_cann.RData (22.7 KB)
Code to run the model is below
load(file = "amphUseInOTP_cann.RData") # object is amphUseInOTP_cann
# give the data object a generic name. Note the days' cannabis use variable is called `outcome` abd is a bounded count between 0 and 28
workingDF <- amphUseInOTP_cann
workingDF %>%
distinct(encID) %>%
nrow -> nEnc
# run betabinomial spline model
fit_betabinomial_cann_s2_ps <- brm(formula = outcome | trials(set) ~ ats_baseline + s(yearsFromStart) + s(yearsFromStart, ats_baseline) + (1 | encID),
data = workingDF,
family = beta_binomial(),
prior = c(prior(prior = normal(0,4),
class = "b",
coef = "ats_baseline",
tag = "b"),
prior(prior = normal(0,4),
class = "b",
coef = "syearsFromStart_1",
tag = "b"),
prior(prior = normal(0,4),
class = "b",
coef = "syearsFromStartats_baseline_1",
tag = "b"),
prior(prior = normal(0,4),
class = "b",
coef = "syearsFromStartats_baseline_2",
tag = "b"),
prior(prior = normal(0,10),
class = "sds",
coef = "s(yearsFromStart)",
tag = "sds"),
prior(prior = normal(0,10),
class = "sds",
coef = "s(yearsFromStart, ats_baseline)",
tag = "sds")),
save_pars = save_pars(all=TRUE),
seed = 1234,
refresh = 0,
chains = 4,
warmup = 1e3,
iter = 2e3)
# define a logscore function
logscore <- function(x) {
as.matrix(rowSums(log_lik(x)))
}
Now we follow the procedure in the vignette. We bind the draws of the predictions and measures to the posterior draws using posterior::bind_draws(). For this we will use the predictions_as_draws helper function provided by priorsense which transforms predictions from brms to draws objects. The idea is that we are attaching predictions at various levels of the outcome to the posterior draws object, and then doing power-scaling on THOSE predictions to assess prior sensitivity.
First we get the grid of predictions (using the data_grid function from modelr), for each of five candidate levels of amphetamine use at baseline - 0 days amphetamine use in the 28 days prior to baseline, 7 days, 14 days, 21 days, and 28 dyas - at each of five timepoints in the first year of treatment: 0 years from start of treatment, 0.25 years, 0.5 years, 0.75 years, 1 year.
tibble(encID=nEnc+1, set=28) %>%
data_grid(yearsFromStart = seq(0,1,by=0.25),
ats_baseline=c(0,7,14,21,28),
encID,
set) -> predGrid
How many combinations of amphetamine use at baseline and yearsFromStart?
nGrid <- nrow(predGrid)
We also need to create names for each of the 25 ats_baseline by yearsFromStart combinations .
expand_grid(paste0("yearsFromStart",seq(0,1,by=0.25)),
paste0("ats_baseline",c(0,7,14,21,28))) %>%
unite(col = "cell") %>%
pull -> cellNames
Now we add the predictions to the posterior draws using bind_draws and predictions_as_draws
post_draws_s2_ps <- as_draws_df(fit_betabinomial_cann_s2_ps) %>%
bind_draws(log_lik_draws(fit_betabinomial_cann_s2_ps)) %>%
bind_draws(predictions_as_draws(x = fit_betabinomial_cann_s2_ps,
predict_fn = bayes_R2,
prediction_names = "R2",
summary = FALSE)) %>%
bind_draws(predictions_as_draws(x = fit_betabinomial_cann_s2_ps,
predict_fn = logscore,
prediction_names = "logscore")) %>%
bind_draws(predictions_as_draws(x = fit_betabinomial_cann_s2_ps,
predict_fn = posterior_epred,
prediction_names = cellNames,
newdata = predGrid,
allow_new_levels = TRUE))
How many variables created in the function above?
nVarPredDraws <- length(variables(as_draws(post_draws_s2_ps)))
The first ten variables are the posterior estimates, and the variables we want to power scale, that we created in the post_draws_s2_ps object immediately above, were pasted onto the end of the posterior draws objects AFTER the R squared and logscore variable (i.e. the last 25 columns). So we want R-squared, logscore, plus all the new prediction columns. We’ll extract their names first
variables(as_draws(post_draws_s2_ps))[c(1:10,(nVarPredDraws-(nGrid+1)):nVarPredDraws)] -> varsToPS
Now to get the powerscale assessment for the coefs and the predictions. The function actually power-scales all priors unless we tell it what we are specifically interested in (specified above in varsToPS object), the fixed effects coefficients, R-squared, logscore, and the predictions
powerscale_sensitivity(x = post_draws_s2_ps,
variable=varsToPS) %>%
tt() %>%
format_tt(num_fmt="decimal") -> powerscale_sens_cann_s2_ps
powerscale_sens_cann_s2_ps
This is great! We see the sensitivity in the earlier model on the posterior estimates
but NOT in R-squared, logscore, or any of the predictions.
I’m new to all of this but I gather this means that the predictions generated by the model are not susceptible to the prior sensitivity evident in the posterior parameter estimates.
Now for the next step in the workflow outlined in the vignette: selectively power-scaling different priors by specifying the corresponding tag in prior_selection. This is where I get stuck
We introduced a prior on the sds term, so let’s try that
powerscale_plot_quantities(
x = post_draws_s2_ps,
div_measure = NULL,
variable = c("R2",
"logscore",
"yearsFromStart0_ats_baseline0",
"yearsFromStart0_ats_baseline7"),
prior_selection = "sds"
)
This throws the error
Error in powerscale_sequence.priorsense_data(psd, lower_alpha = lower_alpha, :
Assertion on 'prior_selection' failed: Must be of type 'numeric' (or 'NULL'), not 'character'.
So it seems like the prior_selection = "sds" argument is the problem and the problem appears to be that it does not like strings. This cannot be though because when I run the code in the vignette, using a string, I get no such error.
Clearly I am doing something wrong.
Any help much appreciated