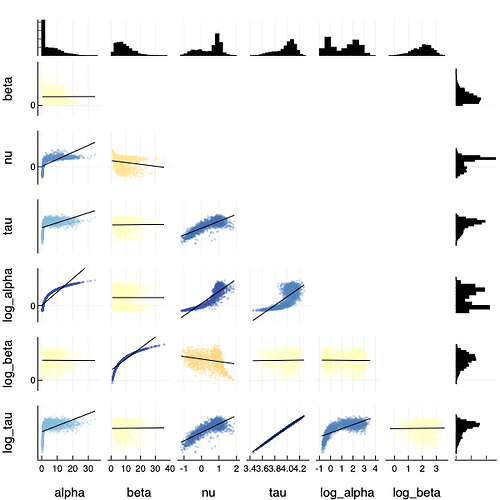
Wow, this is all amazing, thank you for doing such a deep dive into this Mike! That second model is what I was trying to grasp for with the idea of a reparameterisation but I just could not see how to make it work before.
I had to make some tweaks to the second model that you wrote in order for it to sample. Eventually I was able to run this one:
data {
int N;
vector[N] y;
}
parameters {
real<lower=0> alpha;
real<lower=0> epsilon[N];
real nu;
real<lower=0> tau;
}
transformed parameters{
real eps_over_alpha[N];
for(n in 1:N){eps_over_alpha[n] = epsilon[n]*inv(alpha);
}
}
model {
alpha ~ normal(0, 10);
nu ~ normal(0, 10);
tau ~ normal(0, 10);
epsilon ~ exponential(1);
for(n in 1:N){
y[n] ~ normal(nu + eps_over_alpha[n], tau);
}
}
It samples well and fits well for larger values of alpha, but for the smaller values of alpha I tried, the inference isn’t very good. I did a closer look – and I’m not sure what is going wrong here – but both this model and the first function with the improved log(1-phi(x)) function, seem to always return inference of alpha centered around 8.5 no matter what the true alpha is. I’m attaching the code and the two models, the basic one with your updated function and the “betan” one with your cool reparameterisation. Do these fit well on your machine (or did I make an obvious silly mistake in implementing)? Or alternatively does anyone here see the problem? I feel we are so close to solving this problem now, thank you all again for your work on this!
normalhalflaplace-betan-fit.stan (425 Bytes) normalhalflaplace-fit.stan (1.0 KB) single-pareto-lognormal-test.R (5.4 KB)