I apologize in advance for not being able to provide a fully reproducible example, as the model usually takes >24 hours to run and has many thousands of parameters. I hope the information I can provide is sufficient in lieu of that. I’ve read CmdStanR reports error "All variables in all chains must have the same length" after apparently successful sampling - #2 by Robert_Dodier, but it doesn’t solve my issue since I’ve already been redirecting my CSVs to a directory that has 10TB of storage.
I run all of my models on my university’s HPC, which has the following operating system (not sure how little or how much you need, so I included all the info):
- Operating System: Red Hat Enterprise Linux
- CPE OS Name: cpe:/o:redhat:enterprise_linux:7.9:GA:server
- Kernel: Linux 3.10.0-1160.83.1.el7.x86_64
- Architecture: x86-64
I run CmdStan through the CmdStanR interface and R from a custom conda environment, with the below specifications. (Note: I didn’t install CmdStan into a new conda environment as the cmdstan-guide suggests; I have many other R packages in my current conda environment that I need to continue using, and I didn’t want to re-install them in a new environment.)
- R version 4.2.2
- cmdstanr version: 0.5.3
- CmdStan version: 2.31.0
- g++ (GCC) : 8.5.0 20210514 (Red Hat 8.5.0-10)
- GNU Make: 4.3
I recently transitioned to CmdStanR from RStan. When running these same models via RStan, there was always a surge in memory usage once all chains were done running, and the process was creating the stanfit object from all three completed chains. This surge would result in needing 75-100GB of available memory (which I understand, there’s a lot happening in my model) and would result in a stanfit object between 8-14GB (depending on the model). So when running my CmdStan models, I’ve requested similar resources through slurm when I run these models via CmdStanR (so 25-30 cores with 3.15 GB each for two days). I’m sharing all this so you know the computing resources I use when encountering the problems below.
My overall problem is two-fold:
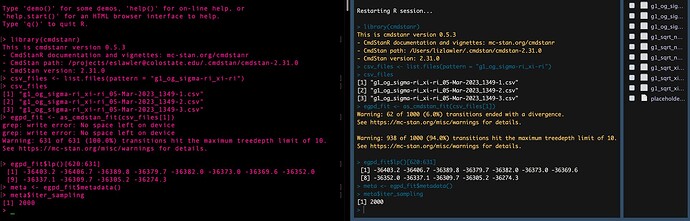
- Once my model completes all three chains successfully (per the console output), for which I’ve been saving CSVs in a specified directory throughout the simulation, I receive two errors:
- “grep: write error: No space left on device” (x3 - it seems this is for each chain?)
- “Error: All variables in all chains must have the same length.”
- Since I’m unable to perform the planned diagnostics on the model object due to the above, I created another R script to recreate the CmdStanMCMC object from the 3 CSVs (one for each chain of the model), then transform that object into an mcmc.list so I can use with the MCMCvis package. But I get the same errors as above (but this time, the grep error happens six times instead of 3).
After continuously receiving the “grep:write error” with all of the models I was running (re problem 1), I reached out to my university’s computing help desk. They determined the issue was likely due to tmp file overflow on the computing node and suggested doing the first two of three things:
- add two lines in your sbatch script: “export TMPDIR=/scratch/alpine/$USER/” and “export TMP=/scratch/alpine/$USER/”
- increase the number of cores in your sbatch script: “#SBATCH --ntasks=”
- command the program/code to auto-clean /tmp directory regularly.
I already included 1 and 2 in the script I created to read in the CSVs and create diagnostic plots (#2 from the original description of my problem). I increased the requested cores to 50, yet I still got the “grep” error and “all variables must have the same length”. I have no idea how to implement 3, and I’m not sure I’d even want to (unless y’all suggest that).
Currently, my biggest concern is being able to create a CmdStanMCMC object from the CSVs of the already fitted chains (for each of the ten models that have already been run). But I still have many other models I need to run, and I’d like to avoid these “grep” and “all variables must be the same length” errors when running models in the future.
Below, I’ve attached the R scripts and console output for both parts of my problem, so you can see more details (although there isn’t much else to say). I’ve included the stan file for one of these models in case that’s helpful. I’ve also included a link with the three CSVs corresponding to this specific model; please note that the zip is 1.65GB, and each file is 2.5GB. Not sure if any or all of this information will help diagnose my problem.
Model run and output:
g1_sigma-ri_xi-ri.stan (8.9 KB)
g1_NUTS_sampling.R (3.0 KB)
g1_og_sigma-ri_xi-ri_894542.txt (10.4 KB)
zip_of_csvs
Diagnostic plot script and output:
g1_dx_plots.R (2.1 KB)
g1_dxplots_og_sigma-ri_xi-ri_925896.txt (1.2 KB)
Thank you in advance for your help and for taking the time to read this! I really appreciate it.
Liz (she/her)